PeptideAtlas: Home Overview Contacts Publications Software Database Schema Feedback Funding FAQ
Related: SRMAtlas PASSEL SWATHAtlas
Spectral Libs: Libraries + Info SpectraST Search
Glossary/Terms: Atlas nomenclature Protein ID terms
LOG IN

The long term goal of the PeptideAtlas project is full annotation of eukaryotic genomes through a thorough validation of expressed proteins. The PeptideAtlas provides a method and a framework to accommodate proteome information coming from high-throughput proteomics technologies. The online database administers experimental data in the public domain. We encourage you to contribute to the database.
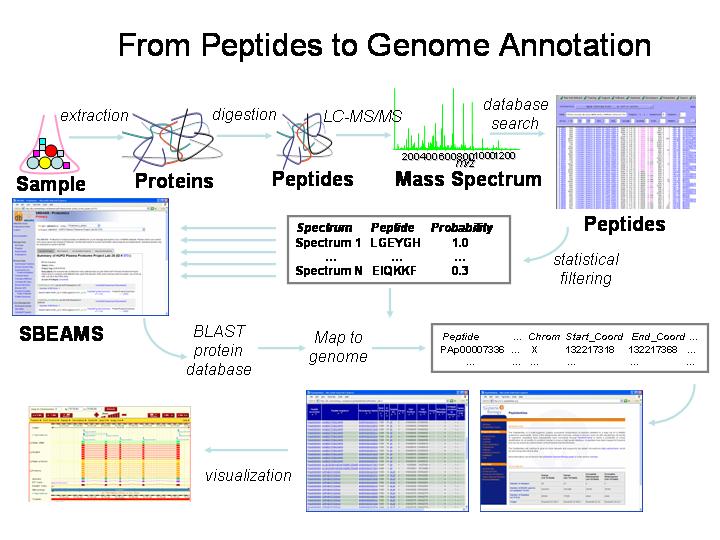
Details of the PeptideAtlas construction can be found within the first publication. Briefly, the general outline of obtaining high quality peptide sequences, mapping, and storing in a database is shown in the figure below and outlined here.
► A protein mixture sample is prepared (perhaps labeled, digested with trypsin, purified, separated using chromotography).
► The sample is run through a mass spectrometer (e.g., ESI MS/MS).
► The MS/MS spectra are compared to theoretical spectra (Comet, X!Tandem) or actual spectra (SpectraST) to identify possible peptides.
► The peptide identifications are scored, formed into false and true positive distributions, and subsequently filtered to retain only the highest scoring identifications (PeptideProphet).
► The peptide sequences are compared to protein sequence databases (e.g. for human, we use the Ensembl, IPI, and Swiss-Prot protein sequence databases). As the peptides are identified in a given protein, so are their locations relative to the protein start (CDS coordinates).
► The peptide locations in chromosomal coordinates are calculated from the CDS coordinates.
► The protein identifications are clustered and annotated (ProteinProphet).
► The data are stored in the SBEAMS database and can be accessed through web pages (see Browse Peptides). The peptides are assigned a unique identity of the form PAp[8-digit number], such as PAp0000001 in our database, but can also be found via their sequences.
The PeptideAtlas is stored using a database schema which accommodates different builds of PeptideAtlas, different versions of ENSEMBL, different organisms (for example, human, fly, mouse), and different reference protein sequence sets as starting material.